Unique microenvironment microarrays give new results
Saturday, 28 February 2009
Medical researchers are praising adult stem cells and their more committed kin, progenitor cells, for their ability to produce different types of specialized cells. The potential of using these cells to repair or replace damaged tissue holds great promise for cancer therapies and regenerative medicine. However, the question that must first be answered is what determines the ultimate fate of a stem or progenitor cell? A team of researchers led by Berkeley Lab's Mark LaBarge and Mina Bissell appear to be well on the road to finding out.
 Working with unique microenvironment microarrays (MEArrays) of their own creation, LaBarge, Bissell, and their collaborators have shown that the ultimate fate of a stem or progenitor cell in a woman's breast – whether the cell develops normally or whether it turns cancerous – may depend upon signals from multiple microenvironments.
"We found that adult human mammary stem and progenitor cells exhibit impressive plasticity in response to hundreds of unique combinatorial microenvironments," said LaBarge, a cell and molecular biologist in Berkeley Lab's Life Sciences Division.
"Our results further suggest that rational modulation of the microenvironmental milieu can impose specific differentiation phenotypes on normal stem or progenitor cells, and perhaps even impose phenotypically normal behaviour on malignant cells during tissue genesis. All of this points to the rational manipulation of adult stem and progenitor cells as a promising pathway for beneficial therapies."
Previous studies on how microenvironments affect the development of adult human stem or progenitor cells have been based on the behaviour of these cells in culture (in vitro) where they are exposed to a single molecular agent. However, when these cells are in an actual human being (in vivo) they are surrounded by a multitude of other cells plus a supporting network of fibrous and globular proteins called the extracellular matrix (ECM), as well as many other nearby molecules, all of which may be simultaneously sending them instructional signals.
"With our MEArrays, we can use combinations of proteins from a select tissue to create multiple microenvironments on a single chip about two square centimetres in area," said LaBarge.
"We think this approach will give us a much more realistic picture as to how stem and progenitor cells actually behave in vivo."
"We have demonstrated that each discrete cell fate decision requires the integration of multiple pathways, and we have identified combinations of components in the human mammary microenvironment that impose distinct cell fates. These results are exciting because they indicate that we can test a large number of effectors and determine which ones to use to direct the fate of adult stem and progenitor cells. This give hope that one day - sooner rather than later - the information could be used for therapy," said Bissell, a Distinguished Scientist with Berkeley Lab's Life Sciences Division and one of the world's leading researchers on breast cancer.
Collaborating with LaBarge and Bissell on this study were Jason Ruth, now at the University of Pennsylvania, Martha Stampfer of Berkeley Lab, Celeste Nelson, now with Princeton University, and Rene Villadsen, Agla Fridriksdottir and Ole Petersen, of the Panum Institute in Denmark.
Human breast tissue harbours two types of epithelial cells: luminal - the cells that are able to produce milk and generally the ones that become cancerous; and myoepithelial – the cells that surround the luminal cells and push milk down the ducts to the nipples, but which rarely become cancerous. Like cells in other types of tissue these breast epithelial cells are spawned from stem and progenitor cells that despite being primitive - essentially a cellular blank slate - possess the exact same genome as their differentiated daughters. Once it was widely held that adult stem and progenitor cells intrinsically "know" when to self-renew and when to differentiate into one specific tissue cell or another based on pre-determined genetic programs. However, pioneering research by Bissell, in which it has been demonstrated that interactions between an epithelial breast cell and its ECM play a major role in determining whether that cell becomes cancerous, pointed the way to the idea that the ultimate fate of a stem or progenitor cell is heavily influenced by interactions with its neighbouring microenvironments.
Working with unique microenvironment microarrays (MEArrays) of their own creation, LaBarge, Bissell, and their collaborators have shown that the ultimate fate of a stem or progenitor cell in a woman's breast – whether the cell develops normally or whether it turns cancerous – may depend upon signals from multiple microenvironments.
"We found that adult human mammary stem and progenitor cells exhibit impressive plasticity in response to hundreds of unique combinatorial microenvironments," said LaBarge, a cell and molecular biologist in Berkeley Lab's Life Sciences Division.
"Our results further suggest that rational modulation of the microenvironmental milieu can impose specific differentiation phenotypes on normal stem or progenitor cells, and perhaps even impose phenotypically normal behaviour on malignant cells during tissue genesis. All of this points to the rational manipulation of adult stem and progenitor cells as a promising pathway for beneficial therapies."
Previous studies on how microenvironments affect the development of adult human stem or progenitor cells have been based on the behaviour of these cells in culture (in vitro) where they are exposed to a single molecular agent. However, when these cells are in an actual human being (in vivo) they are surrounded by a multitude of other cells plus a supporting network of fibrous and globular proteins called the extracellular matrix (ECM), as well as many other nearby molecules, all of which may be simultaneously sending them instructional signals.
"With our MEArrays, we can use combinations of proteins from a select tissue to create multiple microenvironments on a single chip about two square centimetres in area," said LaBarge.
"We think this approach will give us a much more realistic picture as to how stem and progenitor cells actually behave in vivo."
"We have demonstrated that each discrete cell fate decision requires the integration of multiple pathways, and we have identified combinations of components in the human mammary microenvironment that impose distinct cell fates. These results are exciting because they indicate that we can test a large number of effectors and determine which ones to use to direct the fate of adult stem and progenitor cells. This give hope that one day - sooner rather than later - the information could be used for therapy," said Bissell, a Distinguished Scientist with Berkeley Lab's Life Sciences Division and one of the world's leading researchers on breast cancer.
Collaborating with LaBarge and Bissell on this study were Jason Ruth, now at the University of Pennsylvania, Martha Stampfer of Berkeley Lab, Celeste Nelson, now with Princeton University, and Rene Villadsen, Agla Fridriksdottir and Ole Petersen, of the Panum Institute in Denmark.
Human breast tissue harbours two types of epithelial cells: luminal - the cells that are able to produce milk and generally the ones that become cancerous; and myoepithelial – the cells that surround the luminal cells and push milk down the ducts to the nipples, but which rarely become cancerous. Like cells in other types of tissue these breast epithelial cells are spawned from stem and progenitor cells that despite being primitive - essentially a cellular blank slate - possess the exact same genome as their differentiated daughters. Once it was widely held that adult stem and progenitor cells intrinsically "know" when to self-renew and when to differentiate into one specific tissue cell or another based on pre-determined genetic programs. However, pioneering research by Bissell, in which it has been demonstrated that interactions between an epithelial breast cell and its ECM play a major role in determining whether that cell becomes cancerous, pointed the way to the idea that the ultimate fate of a stem or progenitor cell is heavily influenced by interactions with its neighbouring microenvironments.
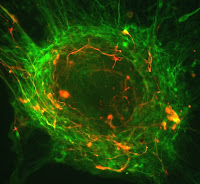
 This is a section of a normal mammary gland that has been stained to find the terminal duct zones that are enriched for mammary stem cells (red) and to visualize a Notch signalling pathway (green). Impairing the Notch pathway caused malignant breast cancer cells to revert to a normal phenotype. Credit: Mark LaBarge.
This is a section of a normal mammary gland that has been stained to find the terminal duct zones that are enriched for mammary stem cells (red) and to visualize a Notch signalling pathway (green). Impairing the Notch pathway caused malignant breast cancer cells to revert to a normal phenotype. Credit: Mark LaBarge.
"Adult stem cells are maintained inside a specialized microenvironment called a niche, whereas progenitor cells migrate to surrounding microenvironments that are distinct from the one around the niche," said LaBarge.
"The ability of adult stem cells to self-maintain, as well as to give rise to progenitor cells that are targeted to become a specific tissue cell, indicates an ability to respond to changing microenvironmental demands, which would mean that a stem or progenitor cell is receiving instructional information from its surroundings."
The fact that normal cells often lose their tissue-specific functions when placed in culture is further evidence of cell fate being tied in to signals from the microenvironment. However, proving such a hypothesis has been difficult in the past because the composition of cell microenvironments is extremely complex and requires a method by which a combination of carefully choreographed interactions can be observed. Given that experiments with human adult stem cell niches cannot be done in vivo and that scientists can only learn so much from mouse models, this means that cell culture studies must be done under as close as possible to in vivo conditions.
"Our technology mimics actual in vivo conditions and enables us to perform highly parallel functional analysis of combinatorial microenvironments, and image analysis of 3-D organotypic cultures and micro patterned culture substrata," said LaBarge.
"The 3-D capability is crucial because our studies show that orientation of the stem or progenitor cells with respect to the signalling molecules can be critical to what happens next."
The MEArrays were fabricated using micro patterning technology originally adapted by co-author Nelson that LaBarge "tweaked." A robot imprinted arrays of 2,304 individual combinations of molecules onto a rubber-coated glass microscope slide (the rubber facilitates adsorption of the proteins onto the slide). An individual MEArray consisted of 192 unique combinatorial microenvironments replicated 12 times, with a plastic barrier running along the perimeter so that cell cultures could be placed on top.
In addition to possible contributors to the stem cell niche, the microenvironments also comprised many ECM and signalling molecules that are expressed in the breast but had not been directly linked to stem cell function before.
In all, adult mammary stem and progenitor cells were exposed to 8,000 different combinations of breast tissue protein and biological molecules. LaBarge, Bissell and their collaborators were able to distinguish between effects resulting from cell interactions with other cells and those resulting from cell interactions with the ECM or other signalling molecules. Both immortalized and primary human breast progenitors were analyzed with the MEArrays and the results were used in conjunction with physiologically relevant 3-D human breast cultures. This approach enabled the research team to identify conditions that induced cells to convert into normal breast cell types as well as conditions that kept the cells in their original, non-specialized state.
One of the most intriguing results in this study was the suggestion that modulation of stem and progenitor cell differentiation pathways might be used to "normalize" malignant breast cells.
"Normal and malignant mammary epithelial cells in 3-D cultures have distinct phenotypes," LaBarge said.
"By impairing a signalling pathway known as Notch, we are able to revert malignant breast cancer cells to a normal phenotype."
In previous studies, Bissell and her group had identified signalling pathways that could cause "phenotypic reversion" of breast cancer cells but this had never been tried before with stem cells.
"The MEArray approach may be able to teach us how to direct stem cell function in a therapeutic setting and possibly to re-program non-stem cells to acquire other stem cell fates," said Bissell.
While the MEArrays in this study were used to study adult stem and progenitor cells in breast tissue, the technique should also be applicable to any of the other 200 different types of tissue cells within other organs, LaBarge said.
.........
ZenMaster
For more on stem cells and cloning, go to CellNEWS at http://cellnews-blog.blogspot.com/ and http://www.geocities.com/giantfideli/index.html